Two days after STRATIF-AI was featured at the BME@LIU-day in Linköping, Sweden, it is featured again in the ISMH conference in Technirghiol, Rumania. In Linköping, we had a shared booth with SUND (the spin-off company that converts the digital twins to eHealth products), and produced a roll-up and a poster. This roll-up has now been transported to Rumania, together with Gunnar Cedersund, to be displayed at a 2h session at the ISMH conference. This happens today May 23, at 17-19, CET, and can be viewed online.


ISMH stands for the International Society of Medical Hydrology and Climatology. In other words, the conference deals with water-based treatments, such as spas (German: kurort), mineral waters, mud baths, as well as temperature-based treatments, such as saunas, ice-baths, etc. These are old traditional treatments, that dates back to ancient Greek, at the least, but there is also a growing scientific interest, with 10 times more papers published per year today, compared to 20 years ago. Still, the ISMH is old, by scientific standards, and the conference celebrated its 125th birthday yesterday. In the picture below, you see Prof, MD Gelu Onose, one of our main collaborators in STRATIF-AI, and also one of the the main organizer of the event, who cut the first piece of the cake, during yesterday’s celebration.

While the official motto of the conference is “Evidence-based balneology”, while there are lots of medical doctors and professors here, and while balneology is an established speciality in medicine in quite a few countries in Europe, it is still a field that lies on the fringes of medicine. For instance, in Sweden, balneology is not an established discipline of medicine, and MDs cannot become specialized in this field, as they can in e.g. cardiology, or hepatology. Therefore, balneology is to some extent similar to other alternative treatments, such as yoga, regular exercise and gym-based personal trainers, health coaches, etc. They exist in society, they work with health, there is some scientific support for the health benefits of these activities, but they are to a large extent not integrated in conventional healthcare. This is where the activities and visions of STRATIF-AI can contribute. Our ambition is to go from a doctor-centric fragmented system, to an integrated patient-centric eco-system system (see picture below). In this eco-system, e.g. personal trainers can take over where nurses leave a patient, following a health conversation (as in our clinical study 2 in STRATIF-AI). And in this eco-system, one can add new actors, such as spas and kurorts, as long as there is evidence that their methods work, and as long as the underlying mechanisms can be added to the digital twin’s models and visualizations.

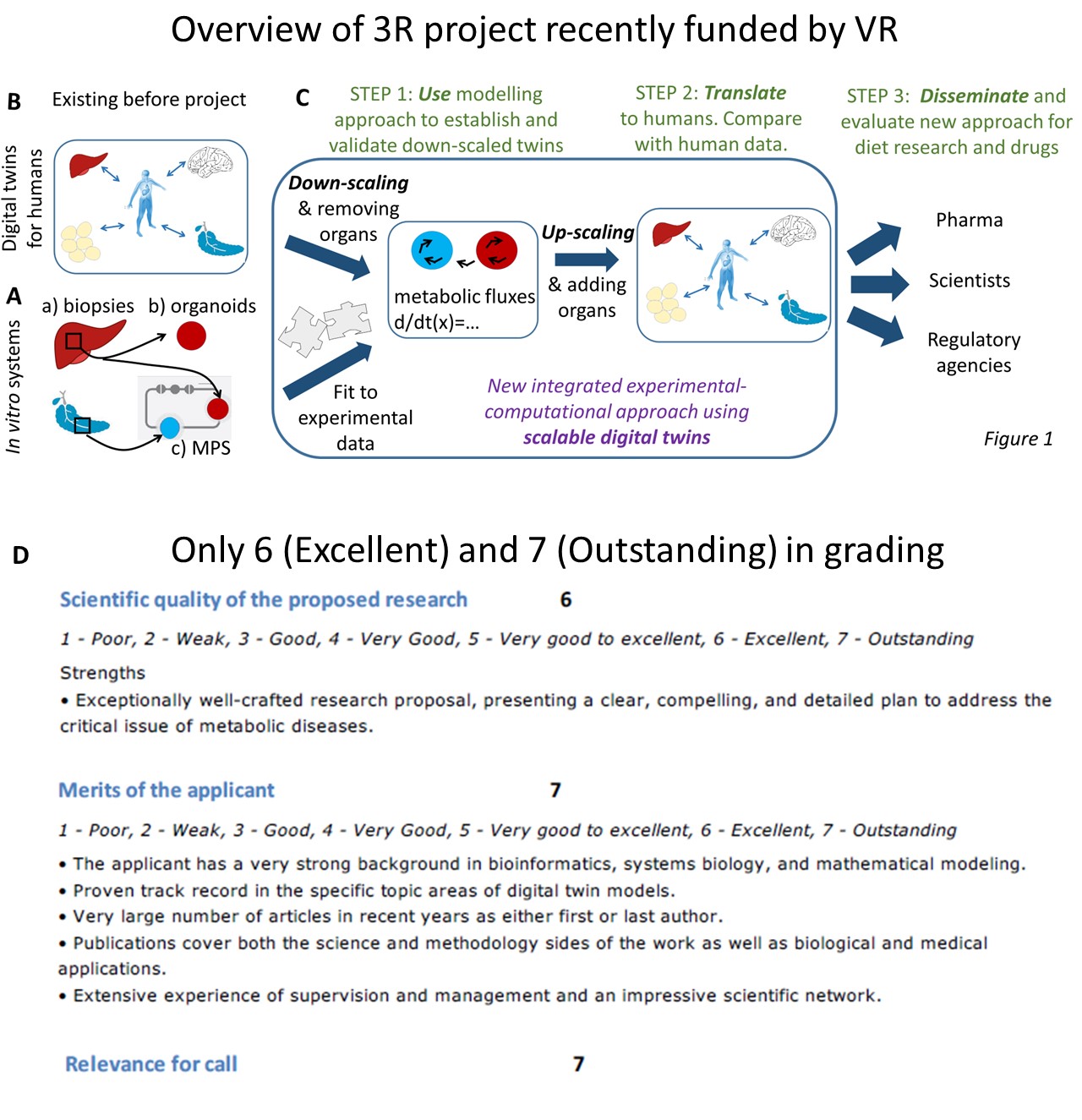
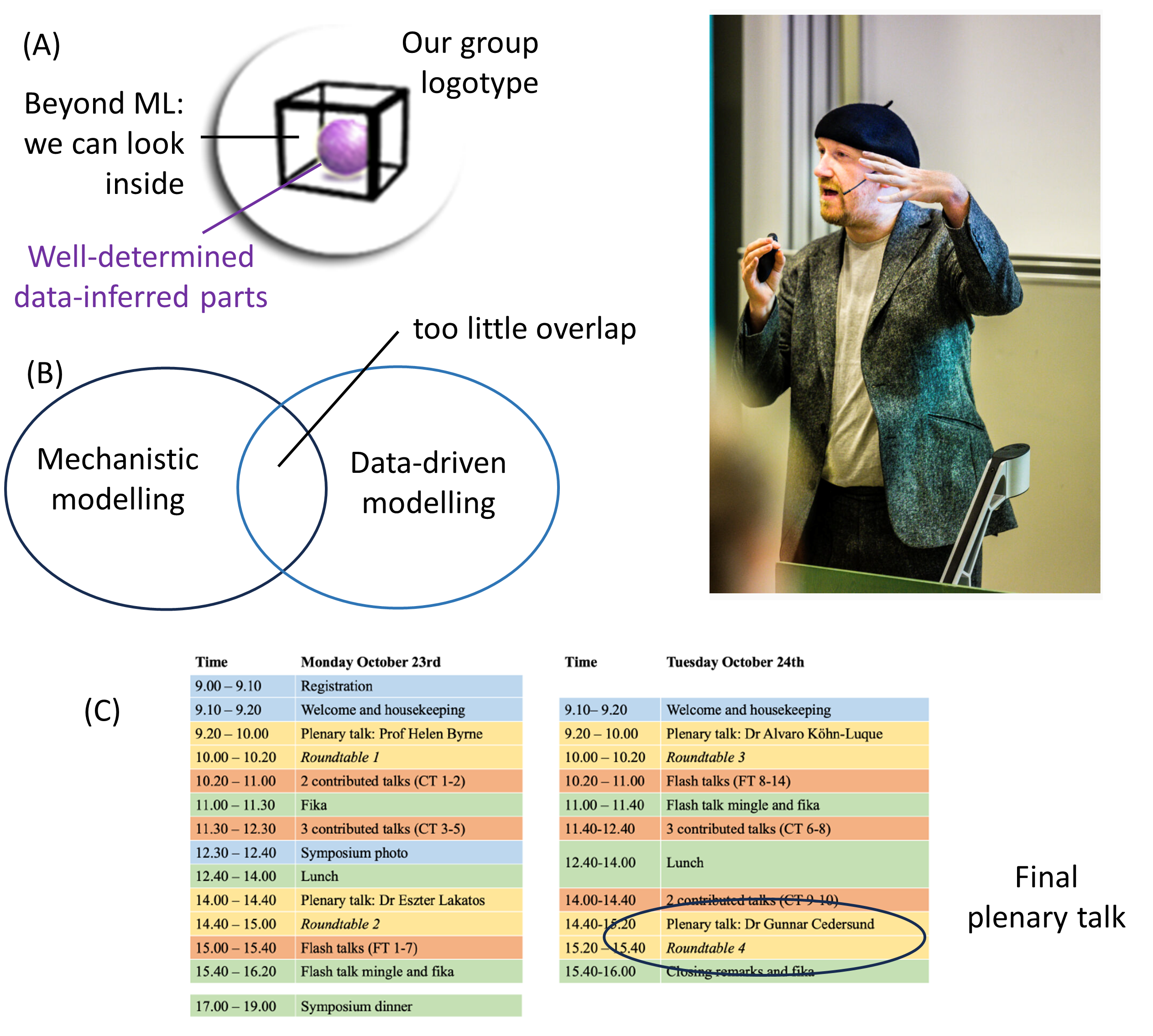
The 2h session we will have is structured as follows: First, Gunnar Cedersund, the coordinator of STRATIF-AI gives an overview of STRATIF-AI, and of the digital twins, which converts knowledge and data to computational models. Thereafter Prof Gelu Onose from SCUBA (in the picture above), and his colleagues Constantin Munteanu and Cristina Popescu (both MDs), give an overview of the scientific basis for prevention and treatment of stroke using different modalities, including balneology. Thereafter, we look at the data integration aspects: Jesper Fellenius from Z2 gives an overview of the personal data vault, Cati Martinez from University of Murcia talks about their work with semantic harmonization and inter-operability, and SIMAVI talk about their work on Electronic Healthcare Records, and app developments, e.g. for rehabilitation of stroke. The session is concluded with a round-table discussion. All things can be followed online, at this link. The general link to the conference is available here.